Crop legumes are an integral part of sustainable agriculture, as several of these species represent an important protein source for both human and animal populations. Legumes engage in a unique and beneficial interaction with a group of soil bacteria, collectively called rhizobia. Rhizobium-legume symbioses lead to the development of new plant organs on the roots called nodules, which host rhizobia. Competent rhizobia within these nodules use a process called nitrogen fixation to convert atmospheric nitrogen, which cannot be used by the plant, into ammonium, which can be used as a nitrogen source. This is a highly valuable process as nitrogen is limited in agricultural systems.
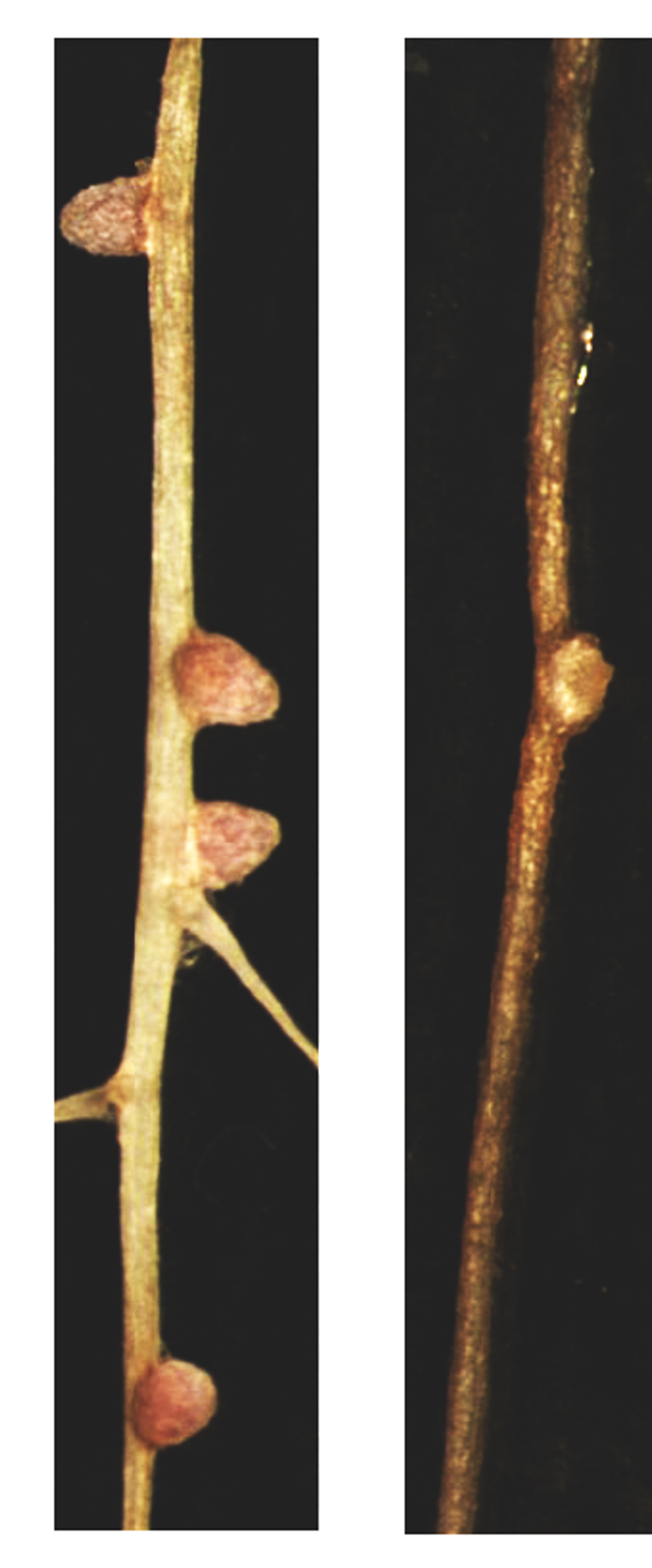
Nitrogen fixation in legumes is highly sensitive to environmental stresses, which limits the exploitation of this process for food production. For instance, we know that soil salinity negatively affects nodulation and inhibits its cognate biochemical process. In a recent study published in the
MPMI journal, Jeanne Harris (The University of Vermont) and her team explored the effects of salinity on barrelclover-rhizobia interaction.
“Since we were interested in learning the underlying mechanism of this interaction, we profiled the plant gene expression in the presence of salt stress, and/or rhizobium, or in the absence of both. Given the inhibitory role of salinity on nodulation, we had expected to see an overall repressive effect on the expression of symbiotic genes under salinity. To our surprise, we found that several genes with well-characterized functions in nodulation are highly induced under salt stress,” explained Sanhita Chakraborty, first author of this study.
“This means that the salt stress actually makes the plant hypersensitive to the signal from the bacterium. This is exciting because it involves an environmental signal that primes the plant response to a biotic signal, a field that remains vastly unexplored in plant sciences,” added Harris.
Through their approach, the researchers were able to identify a previously undescribed set of novel stress-specific rhizobium-responsive genes. “We specifically set out to ask whether the regulation of additional genes might be required under stress conditions that we would never identify under the ideal conditions that we use in the lab. We were excited to find a large group of genes that were specifically induced or repressed only when plants experienced both the salt pretreatment and inoculation with rhizobia,” said Harris.
“Our study provides a set of genes that could be manipulated and introduced into a plant for increased nodulation under salt stress. One or more of these genes could be required for/or prevent nodulation in a stressed environment. Their over-expression or mutation could potentially benefit the agricultural industry,” pointed out Chakraborty.
